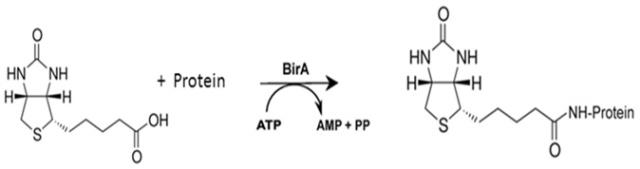
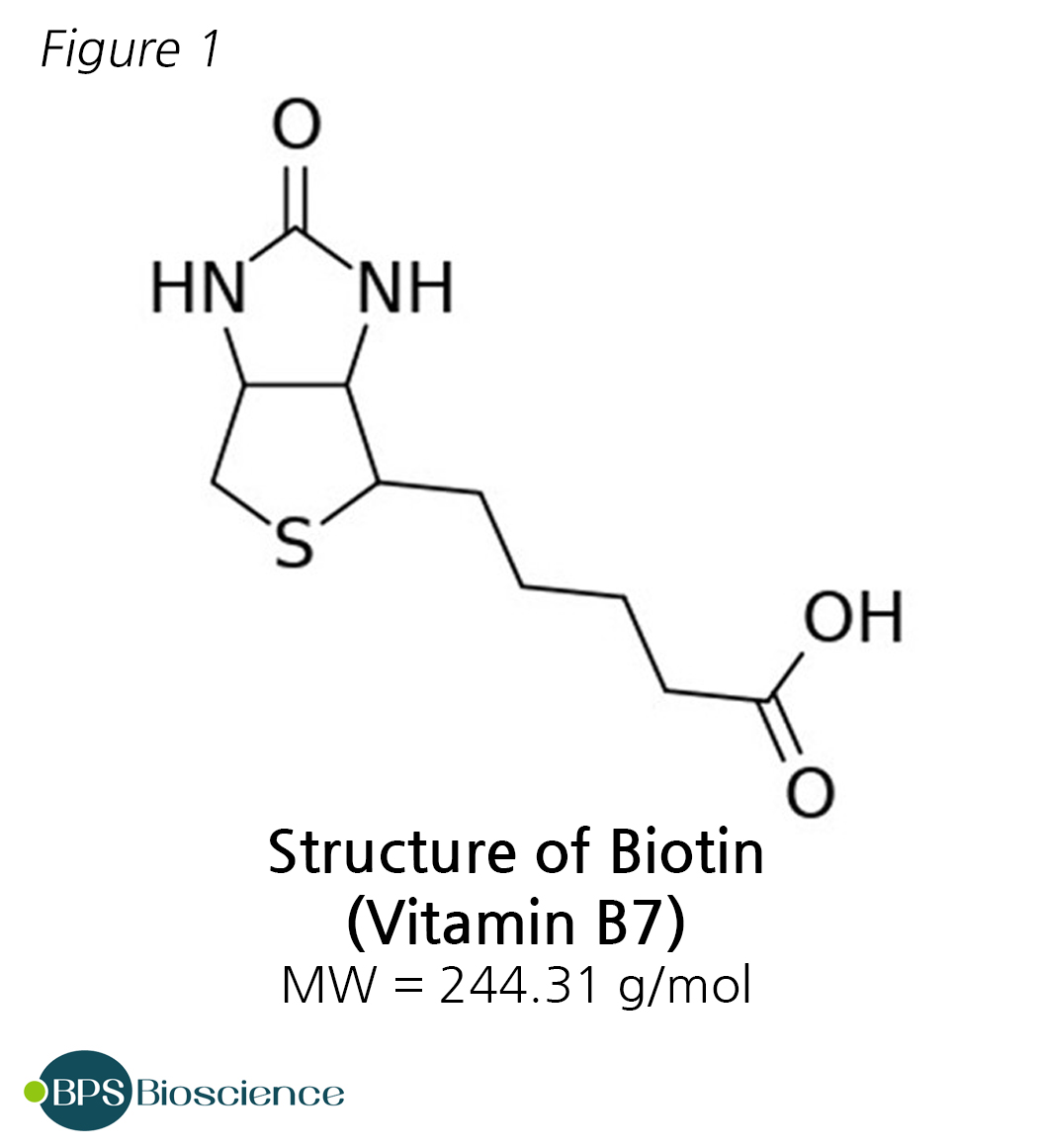
Imaging proteins in live mammalian cells with biotin ligase and monovalent streptavidin. - Abstract - Europe PMC
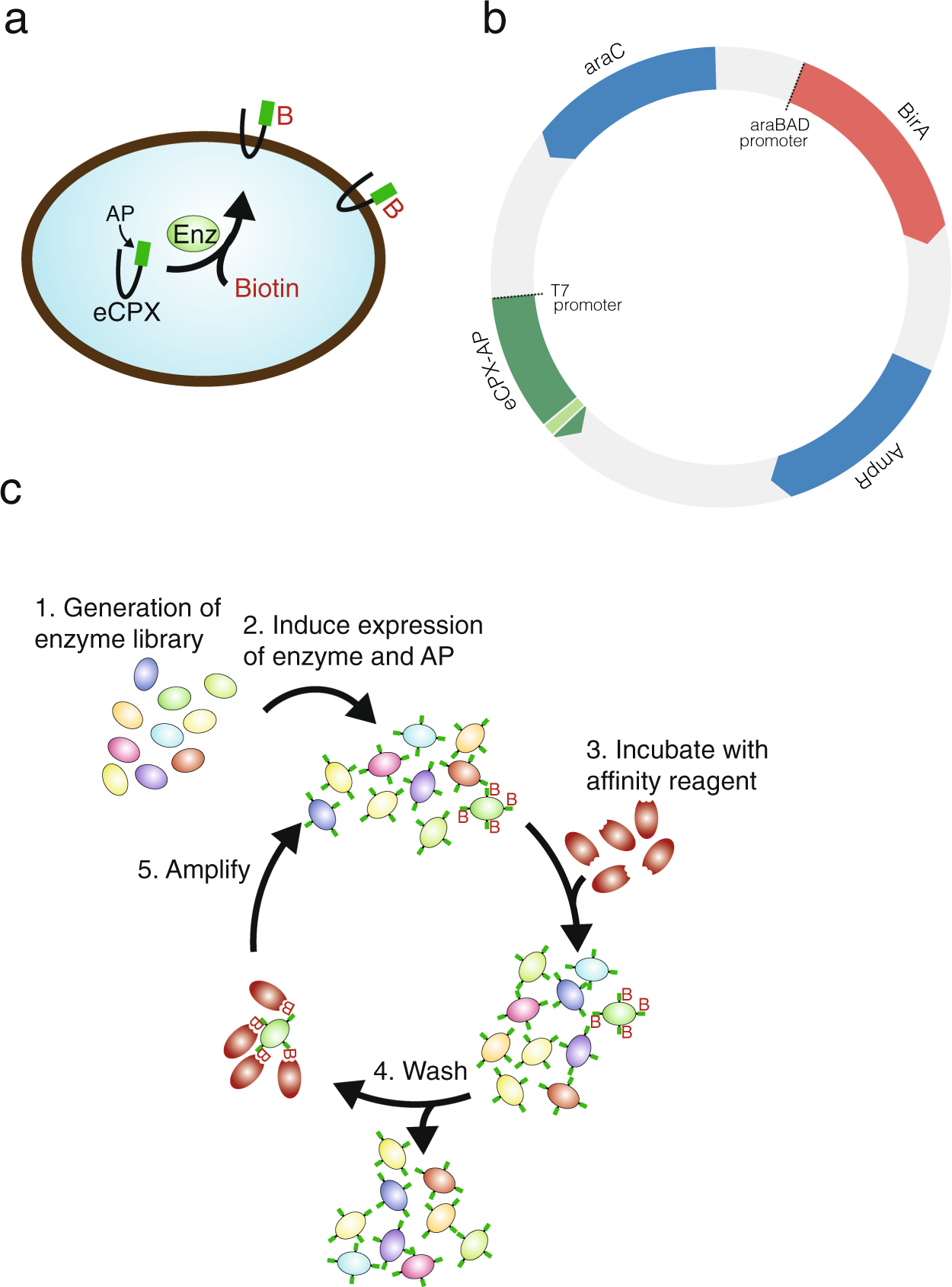
A bacterial display system for effective selection of protein-biotin ligase BirA variants with novel peptide specificity | Scientific Reports

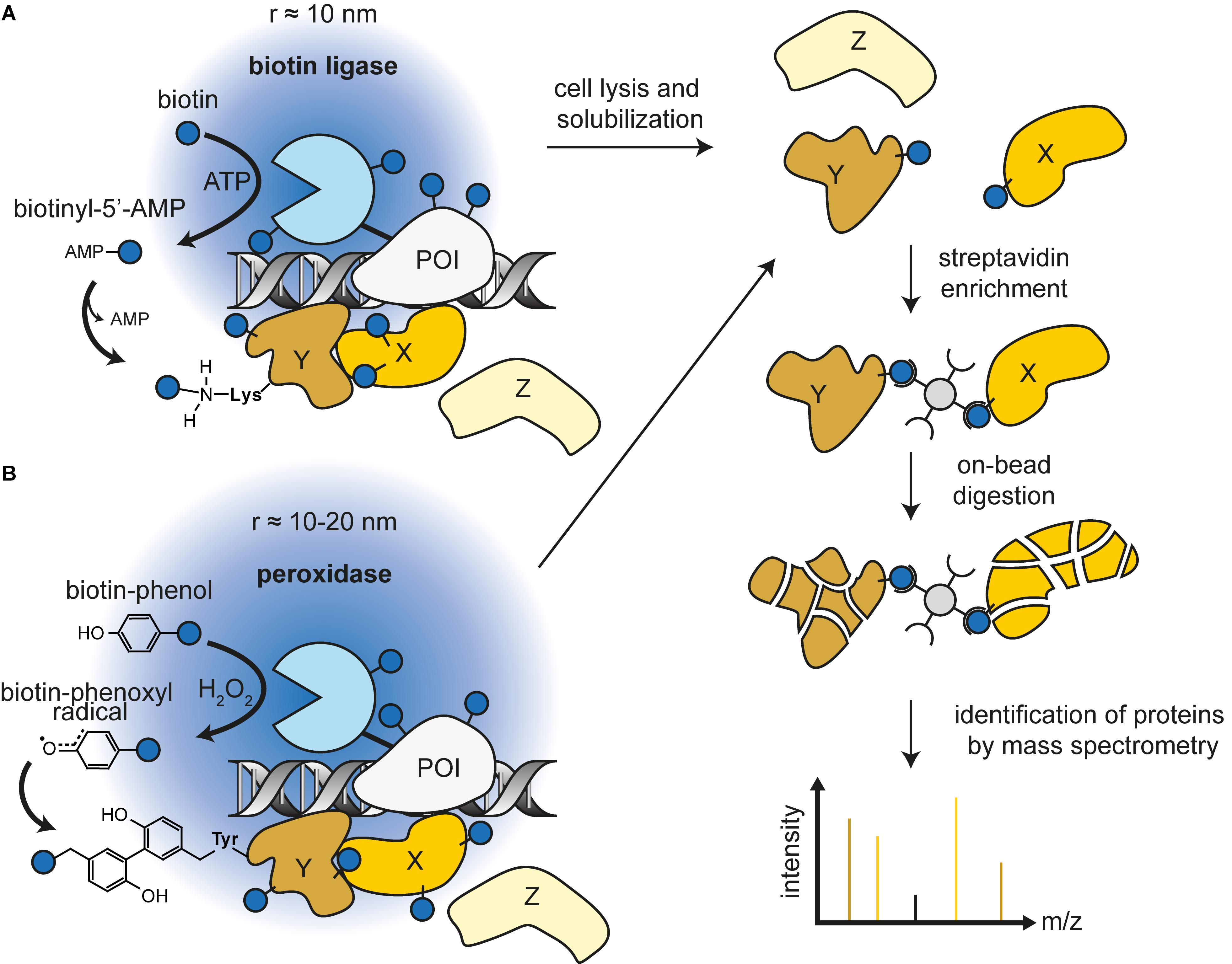
Protein labelling by means of (A) biotin ligase (BirA), (B) lipoic acid... | Download Scientific Diagram

The Atypical Occurrence of Two Biotin Protein Ligases in Francisella novicida Is Due to Distinct Roles in Virulence and Biotin Metabolism | mBio

Overexpression of biotin synthase and biotin ligase is required for efficient generation of sulfur-35 labeled biotin in E. coli | BMC Biotechnology | Full Text

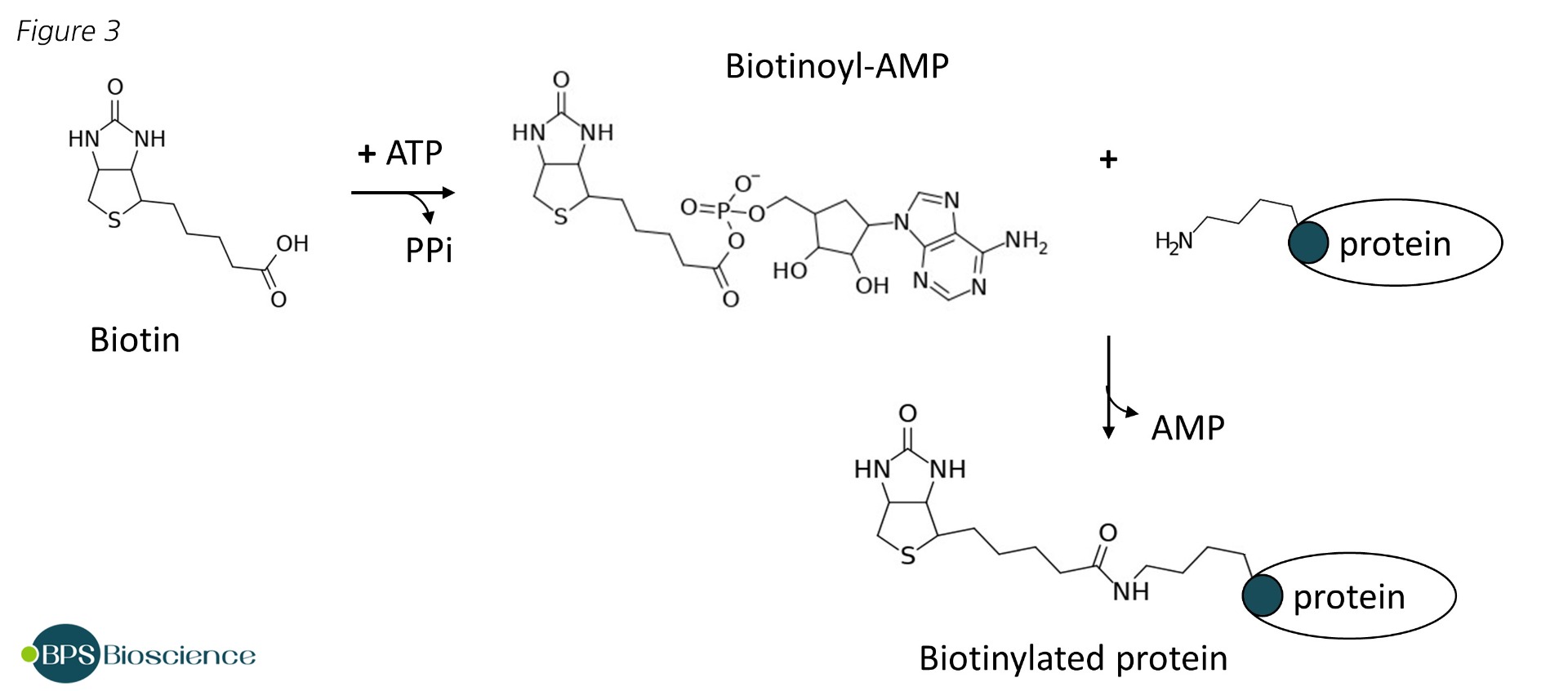
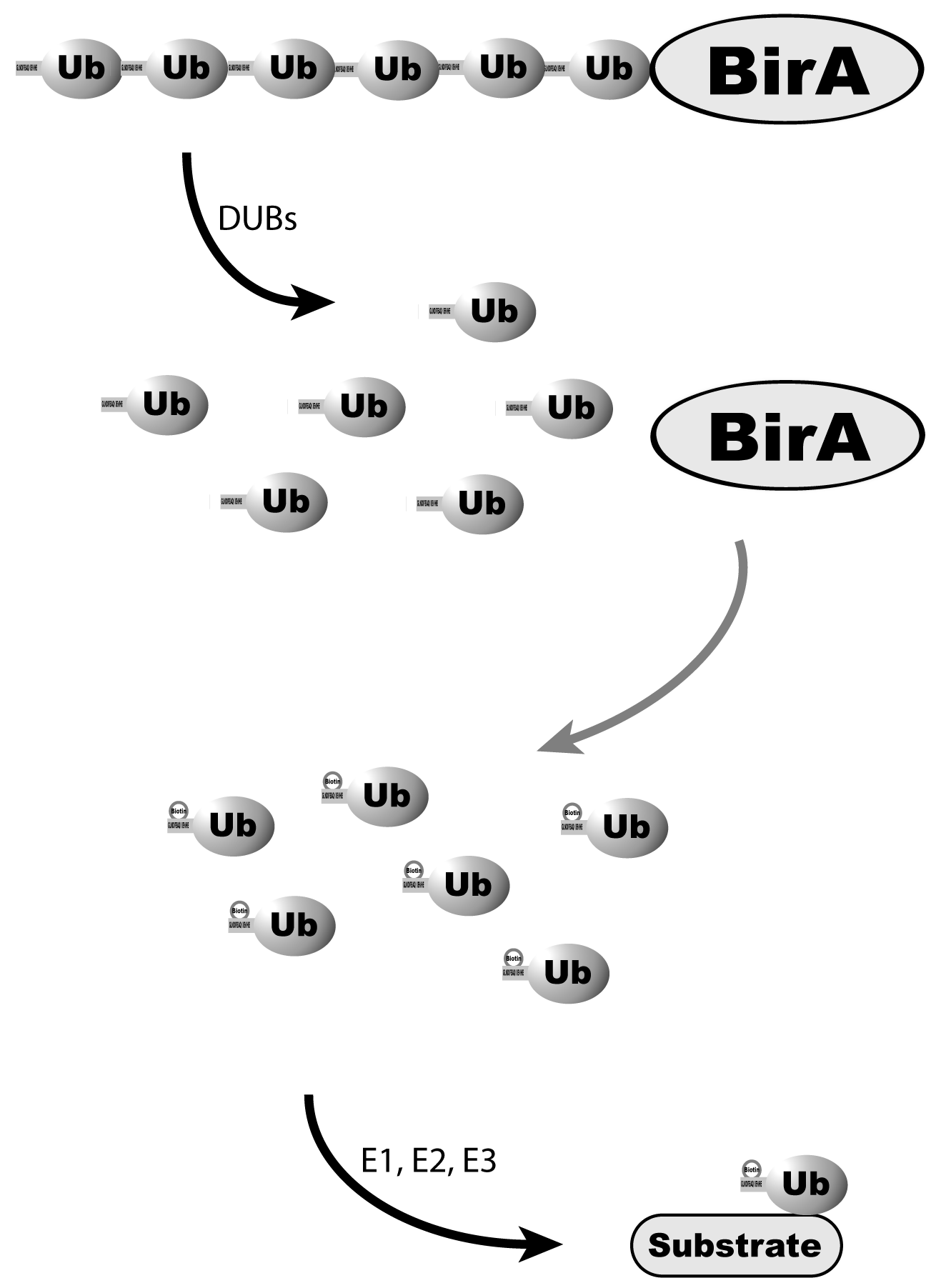
BirA Biotin Protein Ligase for Tagging Proteins and Antibodies – Amid Biosciences | Protein Engineering Company

Bisubstrate Adenylation Inhibitors of Biotin Protein Ligase from Mycobacterium tuberculosis - ScienceDirect

Sulfonamide-Based Inhibitors of Biotin Protein Ligase as New Antibiotic Leads | ACS Chemical Biology
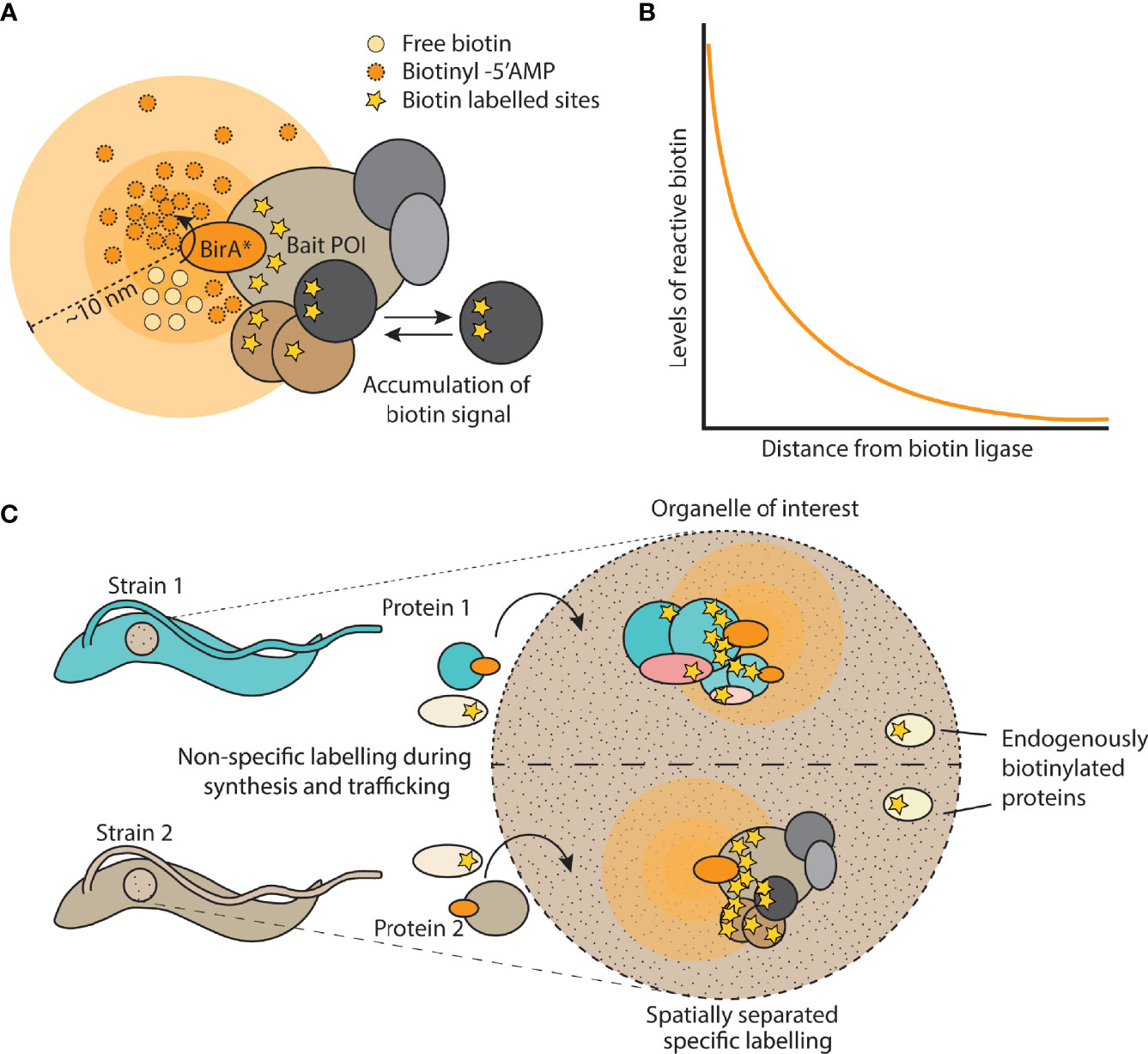
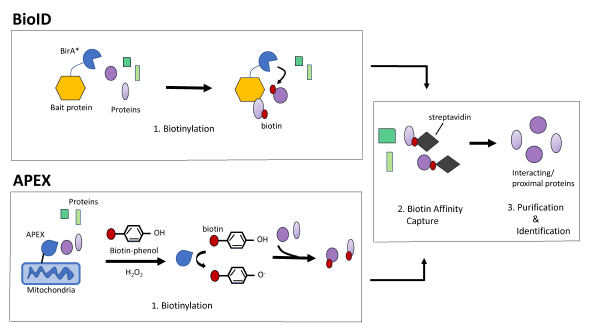
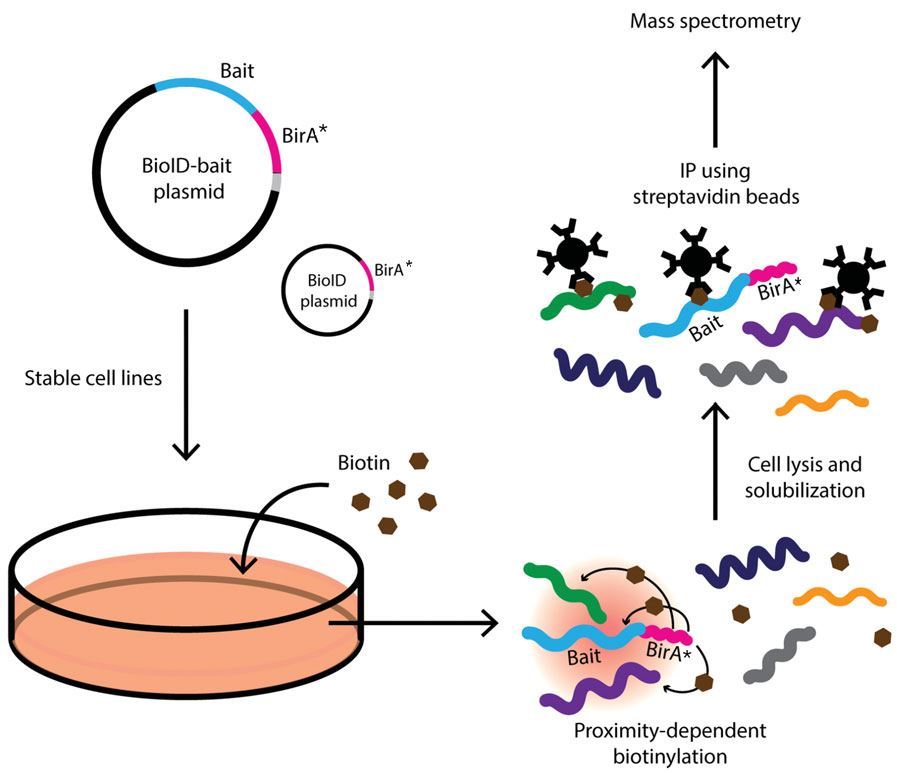
Frontiers | Tag Thy Neighbour: Nanometre-Scale Insights Into Kinetoplastid Parasites With Proximity Dependent Biotinylation